- Motivation
- Variable inclusion plots
- Model stability plots
- Adaptive fence method
- Bootstrapping the lasso
- The
mplotpackage - Diabetes data example
- Future work
September 2015
Overview
Current state of variable selection
- 2 million pages with "model selection"
- 1/2 million pages with "variable selection"
Google Scholar
- 620,000 articles with "model selection"
- 170,000 articles with "variable selection"
Do we really need more?
Current approaches to variable selection
Information Criterion
- AIC
- BIC
- HQIC
Stepwise procedures
- Mallows \(C_p\)
- Sequential testing
Regularisation methods
- Lasso
- SCAD
- LARS (stagewise)
Artificial example

Artificial example
require(mplot) lm.art = lm(y~., data=artificialeg)
| Estimate | Std. Error | t value | Pr(>|t|) | |
|---|---|---|---|---|
| (Intercept) | -0.10 | 0.33 | -0.31 | 0.76 |
| x1 | 0.64 | 0.69 | 0.92 | 0.36 |
| x2 | 0.26 | 0.62 | 0.42 | 0.68 |
| x3 | -0.51 | 1.24 | -0.41 | 0.68 |
| x4 | -0.30 | 0.25 | -1.18 | 0.24 |
| x5 | 0.36 | 0.60 | 0.59 | 0.56 |
| x6 | -0.54 | 0.96 | -0.56 | 0.58 |
| x7 | -0.43 | 0.63 | -0.68 | 0.50 |
| x8 | 0.15 | 0.62 | 0.24 | 0.81 |
| x9 | 0.40 | 0.64 | 0.63 | 0.53 |
Artificial example – Stepwise
step(lm.art)
## Start: AIC=79.3 ## y ~ x1 + x2 + x3 + x4 + x5 + x6 + x7 + x8 + x9 ## ## Df Sum of Sq RSS AIC ## - x8 1 0.2423 163.94 77.374 ## - x3 1 0.6946 164.39 77.512 ## - x2 1 0.7107 164.41 77.517 ## - x6 1 1.3051 165.00 77.698 ## - x5 1 1.4425 165.14 77.739 ## - x9 1 1.6065 165.31 77.789 ## - x7 1 1.8835 165.58 77.873 ## - x1 1 3.4999 167.20 78.358 ## - x4 1 5.7367 169.44 79.023 ## <none> 163.70 79.301 ## ## Step: AIC=77.37 ## y ~ x1 + x2 + x3 + x4 + x5 + x6 + x7 + x9 ## ...
## ## Call: ## lm(formula = y ~ x1 + x2 + x3 + x4 + x5 + x6 + x7 + x9, data = artificialeg) ## ## Coefficients: ## (Intercept) x1 x2 x3 x4 ## -0.1143 0.8019 0.4011 -0.8083 -0.3514 ## x5 x6 x7 x9 ## 0.4927 -0.7738 -0.5772 0.5478
Artificial example – Stepwise
| Estimate | Std. Error | t value | Pr(>|t|) | |
|---|---|---|---|---|
| (Intercept) | -0.11 | 0.32 | -0.36 | 0.72 |
| x1 | 0.80 | 0.19 | 4.13 | 0.00 |
| x2 | 0.40 | 0.18 | 2.26 | 0.03 |
| x3 | -0.81 | 0.19 | -4.22 | 0.00 |
| x4 | -0.35 | 0.12 | -2.94 | 0.01 |
| x5 | 0.49 | 0.19 | 2.55 | 0.01 |
| x6 | -0.77 | 0.15 | -5.19 | 0.00 |
| x7 | -0.58 | 0.15 | -3.94 | 0.00 |
| x9 | 0.55 | 0.19 | 2.90 | 0.01 |
- We've dropped
x8. - All the other variables appear to be significant.
Artificial example – Stepwise
The true data generating process is:
\[y = 0.6x_8 + \varepsilon.\]
art.true = lm(y~x8,data=artificialeg)
| Estimate | Std. Error | t value | Pr(>|t|) | |
|---|---|---|---|---|
| (Intercept) | 0.03 | 0.29 | 0.11 | 0.91 |
| x8 | 0.55 | 0.05 | 10.43 | 0.00 |
Artificial example – Stepwise
The true data generating process is:
\[y = 0.6x_8 + \varepsilon.\]
art.true = lm(y~x8,data=artificialeg)
| Estimate | Std. Error | t value | Pr(>|t|) | |
|---|---|---|---|---|
| (Intercept) | 0.03 | 0.29 | 0.11 | 0.91 |
| x8 | 0.55 | 0.05 | 10.43 | 0.00 |
anova(art.true,step.art)
| Res.Df | RSS | Df | Sum of Sq | F | Pr(>F) | |
|---|---|---|---|---|---|---|
| 1 | 48 | 200.97 | ||||
| 2 | 41 | 163.94 | 7 | 37.03 | 1.32 | 0.2643 |
Some notation
- Say we have a full model with an \(n\times p\) design matrix \(\mathbf{X}\).
- Let \(\alpha\) be any subset of \(p_\alpha\) distinct elements from \(\{1,\ldots,p\}\).
- We can define a \(n\times p_\alpha\) submodel with design matrix \(\mathbf{X}_\alpha\) subset from \(\mathbf{X}\) by the elements of \(\alpha\).
- Denote the set of all possible models as \(\mathcal{A}=\{\{1\},\ldots,\alpha_f\}\).
- Coefficent vector \(\beta\) (and similarly \(\beta_\alpha\))
A smörgåsbord of tuning parameters…
Information Criterion
- Generalised IC: \(\text{GIC}(\alpha;\lambda) = -2\times \text{LogLik}(\alpha)+\lambda p_\alpha\)
With important special cases:
- AIC: \(\lambda=2\)
- BIC: \(\lambda = \log(n)\)
- HQIC: \(\lambda= 2\log(\log(n))\)
Regularisation routines
- Lasso: minimises \(-\text{LogLik}(\alpha) +\lambda\ ||\beta_\alpha||_1\)
- Many variants of the Lasso, SCAD,…
A stability based approach
Aim
To provide scientists/researchers/analysts with tools that give them more information about the model selection choices that they are making.
- Concept of model stability independently introduced by Meinshausen and Bühlmann (2010) and Müller and Welsh (2010) for different linear regression situations.
There are many ways to define stability
- E.g. \(\text{GIC}(\alpha;\lambda) = -2\times \text{LogLik}(\alpha)+\lambda p_{\alpha}\) is unstable when \(\hat{\alpha}(\lambda)\) is selected with dimension \(p_{\hat{\alpha}(\lambda)}\) but, for some small \(\delta>0\), \(\hat{\alpha}(\lambda+\delta)\) selected with \(p_{\hat{\alpha}(\lambda+\delta)} < p_{\hat{\alpha}(\lambda)}\)
- Or perhaps the model size stays the same, but \(\hat{\alpha}(\lambda) \neq \hat{\alpha}(\lambda+\delta)\)
Key idea: small changes should have small effects
A stability based approach
Aim
To provide scientists/researchers/analysts with tools that give them more information about the model selection choices that they are making.
Method
- Investigate model selection criterion across a range of penalty parameter values
- exhaustive searches (where feasible)
- bootstrapping to assess selection stability
- interactive graphical tools
Variable inclusion plots
Variable inclusion plots
Aim
To visualise inclusion probabilities as a function of the penalty multiplier \(\lambda\in [0,2\log(n)]\).
Procedure
- Calculate (weighted) bootstrap samples \(b=1,\ldots,B\).
- For each bootstrap sample, at each \(\lambda\) value, find \(\hat{\alpha}_\lambda^{(b)}\in\mathcal{A}\) as the model with smallest \(\text{GIC}(\alpha;\lambda) = -2\times \text{LogLik}(\alpha)+\lambda p_\alpha\).
- The inclusion probability for variable \(x_j\) is estimated as \(\frac{1}{B}\sum_{b=1}^B 1\{j\in \hat{\alpha}_\lambda^{(b)}\}\).
References
- Müller and Welsh (2010) for linear regression models
- Murray, Heritier, and Müller (2013) for generalised linear models
Artificial example – VIP
require(mplot) vis.art = vis(lm.art) plot(vis.art, which = "vip")
Model stability plots
Artificial example – Loss against size
plot(vis.art, which = "lvk", highlight = "x8")
Artificial example – Loss against size
plot(vis.art, which = "lvk", highlight = "x6")
Model stability plots
Aim
To add value to the loss against size plots by choosing a symbol size proportional to a measure of stability.
Procedure
- Calculate (weighted) bootstrap samples \(b=1,\ldots,B\).
- For each bootstrap sample, identify the best model at each dimension.
- Add this information to the loss against size plot using model identifiers that are proportional to the frequency with which a model was identified as being best at each model size.
References
- Murray, Heritier, and Müller (2013) for generalised linear models
Artificial example – Model stability plot
plot(vis.art, which = "boot")
The adaptive fence
The fence
Notation
- Let \(Q(\alpha)\) be a measure of lack of fit
- Specifically we consider \(Q(\alpha)=-2\text{LogLik}(\alpha)\)
Main idea
The fence (Jiang et al. 2008) is based aroung the inequality: \[ Q(\alpha) \leq Q(\alpha_f) + c \]
- A model \(\alpha\) is inside the fence if the inequality holds.
- For any \(c\geq 0\), the full model is always inside the fence.
- Among the set of models that are inside the fence, model(s) with smallest dimension are preferred.
Illustration
Illustration
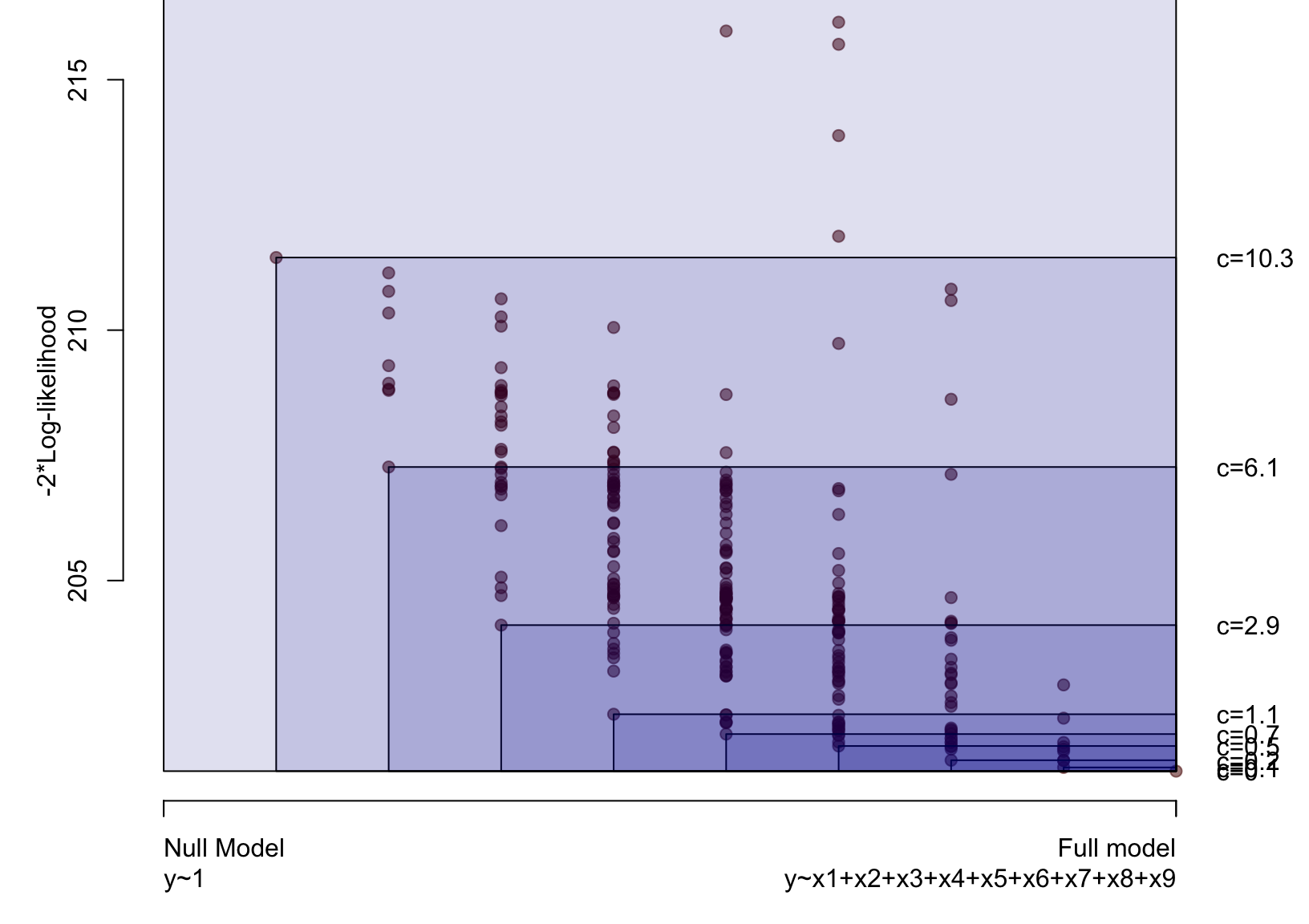
Problem: how to choose \(c\)?
Solution: Bootstrap over a range of values of \(c\).
Procedure
- For each value of \(c\):
- Perform parametric bootstrap under \(\alpha_f\).
- For each bootstrap sample, identify the smallest model that is inside the fence, \(\hat{\alpha}(c)\). Jiang, Nguyen, and Rao (2009) suggest that if there is more than one model, choose the one with the smallest \(Q(\alpha)\).
- Let \(p^∗(\alpha) = P^∗\{\hat{\alpha}(c) = \alpha\}\) be the empirical probability of selecting model \(\alpha\) at a given value of \(c\).
- Calculate \(p^* = \max_{\alpha\in\mathcal{A}} p^*(\alpha)\)
- Plot values of \(p^*\) against \(c\) and find first peak.
- Use this value of \(c\) with the original data.
What does this look like?
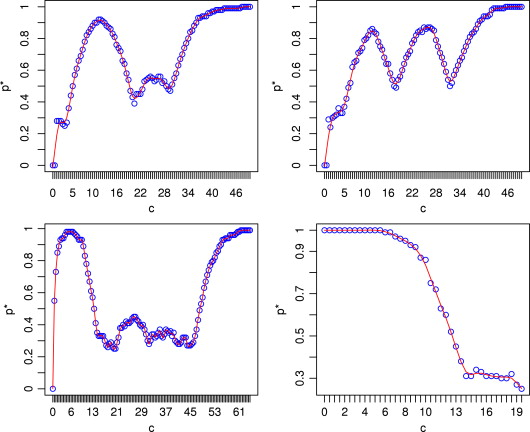
Source: Jiang, Nguyen, and Rao (2009)
Our implementation
Core innovations
- Enhanced the plot of \(p^*\) against \(c\) so that you can see which model is being selected most often
- Allows for consideration of all models that pass the fence at the smallest dimension
- Integration with the
leapsandbestglmpackages - Implemented multicore technology to speed up the embarassingly parallel calculations
Optional innovations
- Intial stepwise procedure to restrict the values of \(c\)
- Can force variables into the model
- Specify the size of the largest model to be considered
- Specify the maximum value for \(c\)
Artificial example – adaptive fence
af.art = af(lm.art, B = 100, n.c = 50, n.cores = 2) plot(af.art)
Speed (linear models; B=50; n.c=25)
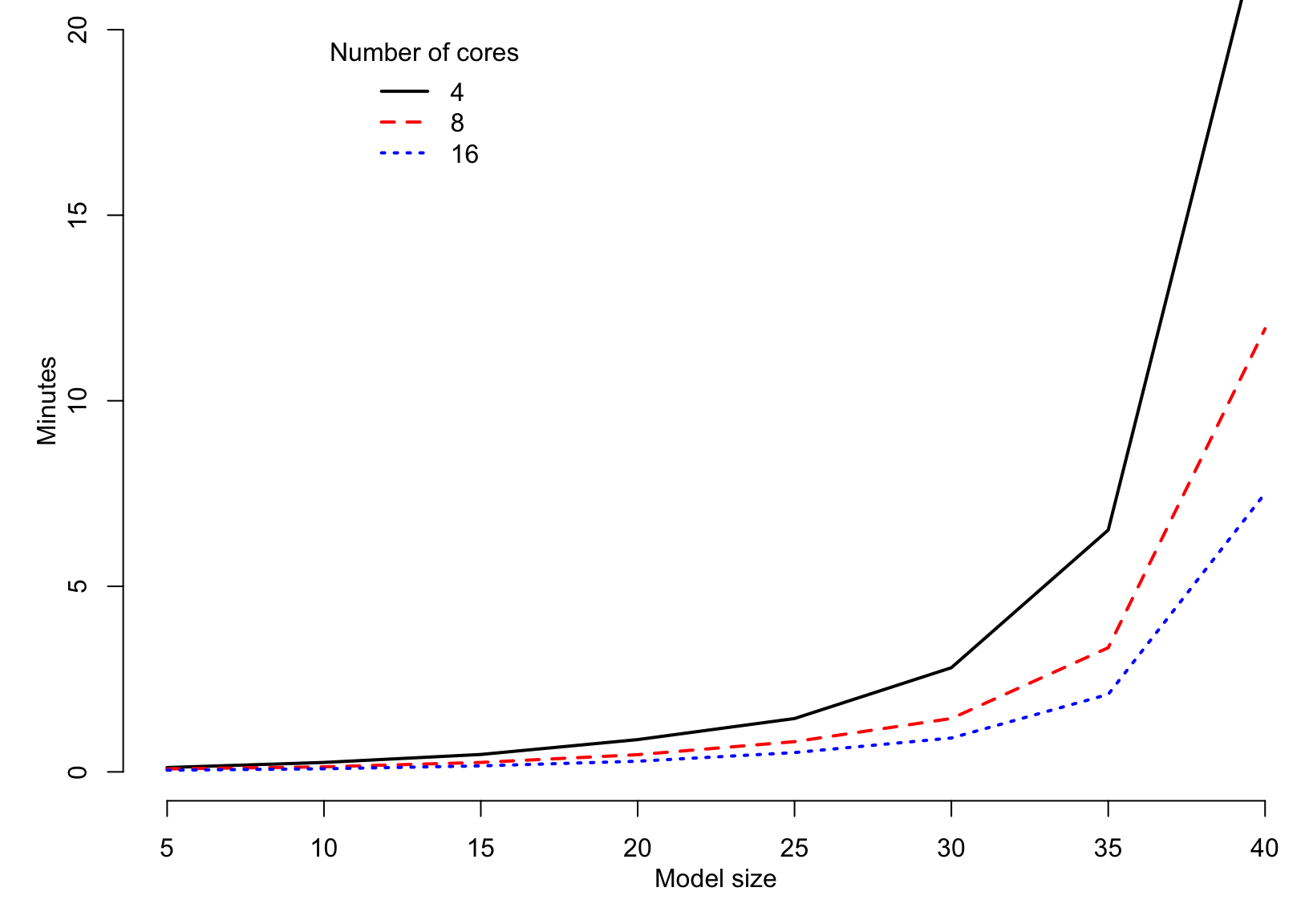
Speed (linear models; B=50; n.c=25)
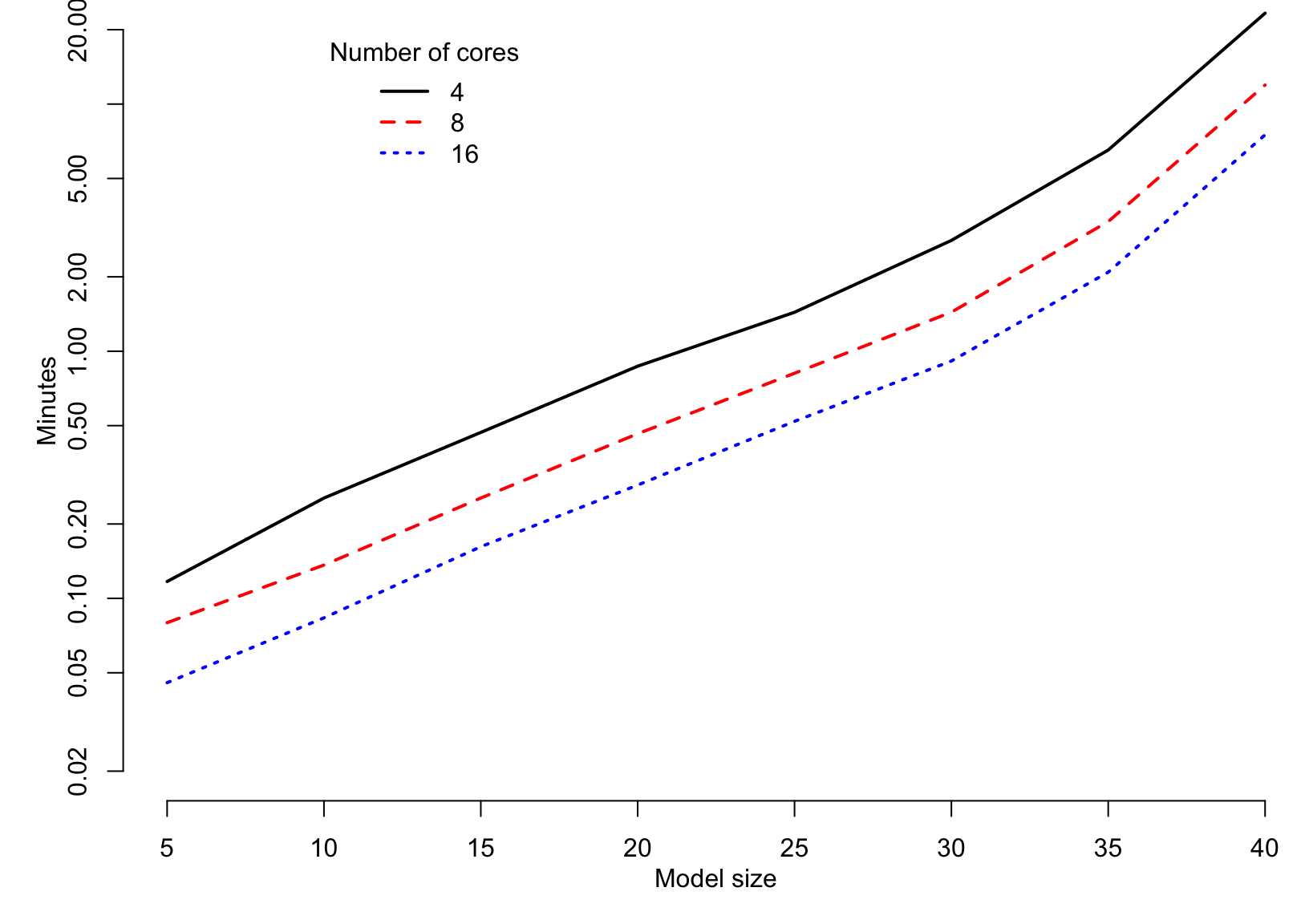
Bootstrapping the lasso
A short history of the lasso
Tibshirani (1996) suggested doing regression with an \(L_1\) norm penalty and called it the lasso (least absolute shrinkage and selection operator).
The lasso parameter estimates are obtained by minimising the residual sum of squares subject to the constraint that \[\sum_j |\beta_j| \leq t.\]
"The encouraging results reported here suggest that absolute value constraints might prove to be useful in a wide variety of statistical estimation problems. Further study is needed to investigate these possibilities." - Tibshirani (1996)
- Implemented easily and quickly through LARS (least angle regression) algorithm of Efron et al. (2004).
Artificial example – Lasso
require(lars) x = as.matrix(subset(artificialeg, select = -y)) y = as.matrix(subset(artificialeg, select = y)) art.lars = lars(x, y) plot(art.lars, xvar = "step", breaks = FALSE, lwd = 2)
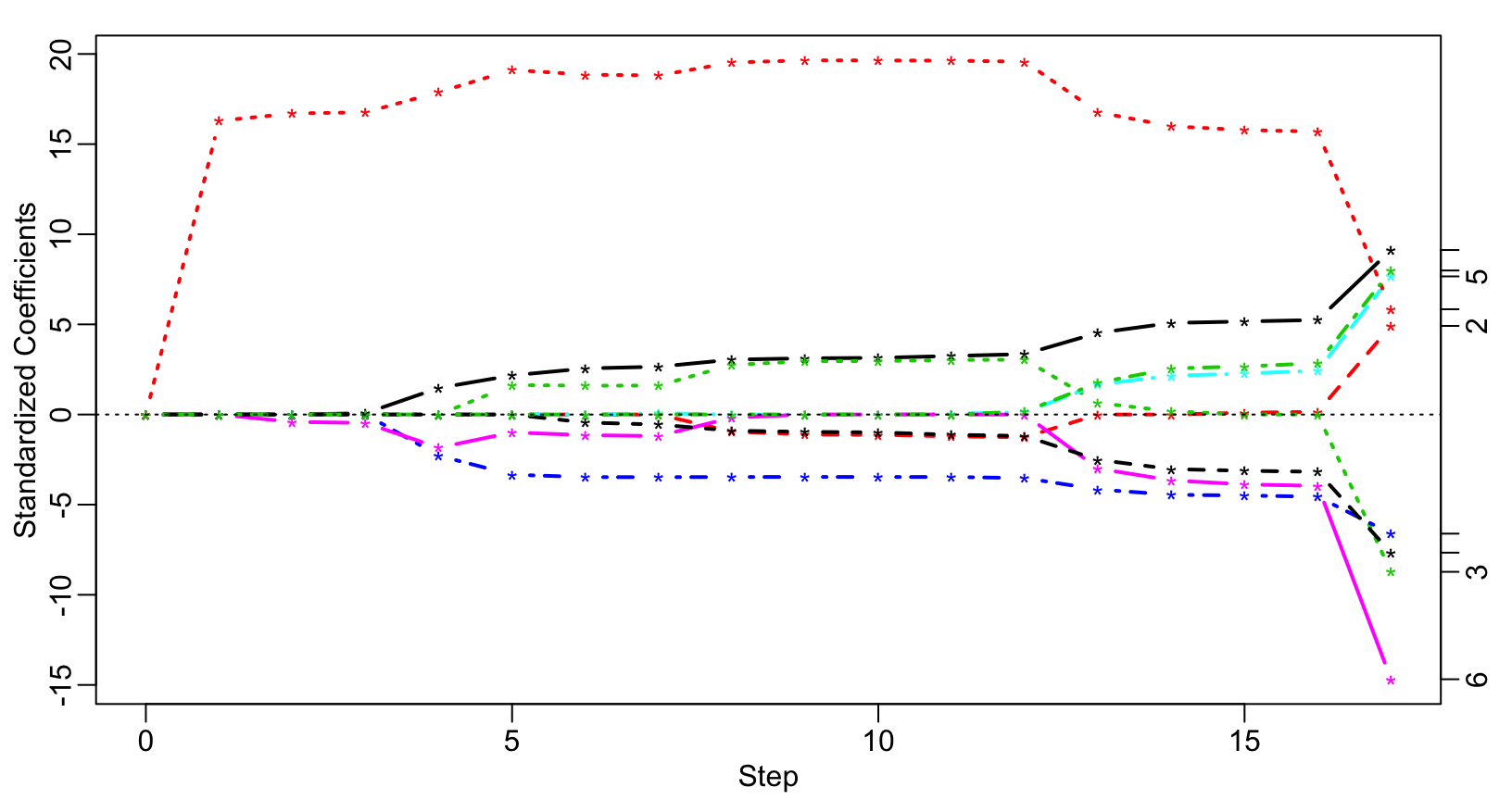
Artificial example – Lasso
plot(art.lars, xvar = "step", plottype = "Cp", lwd = 2)
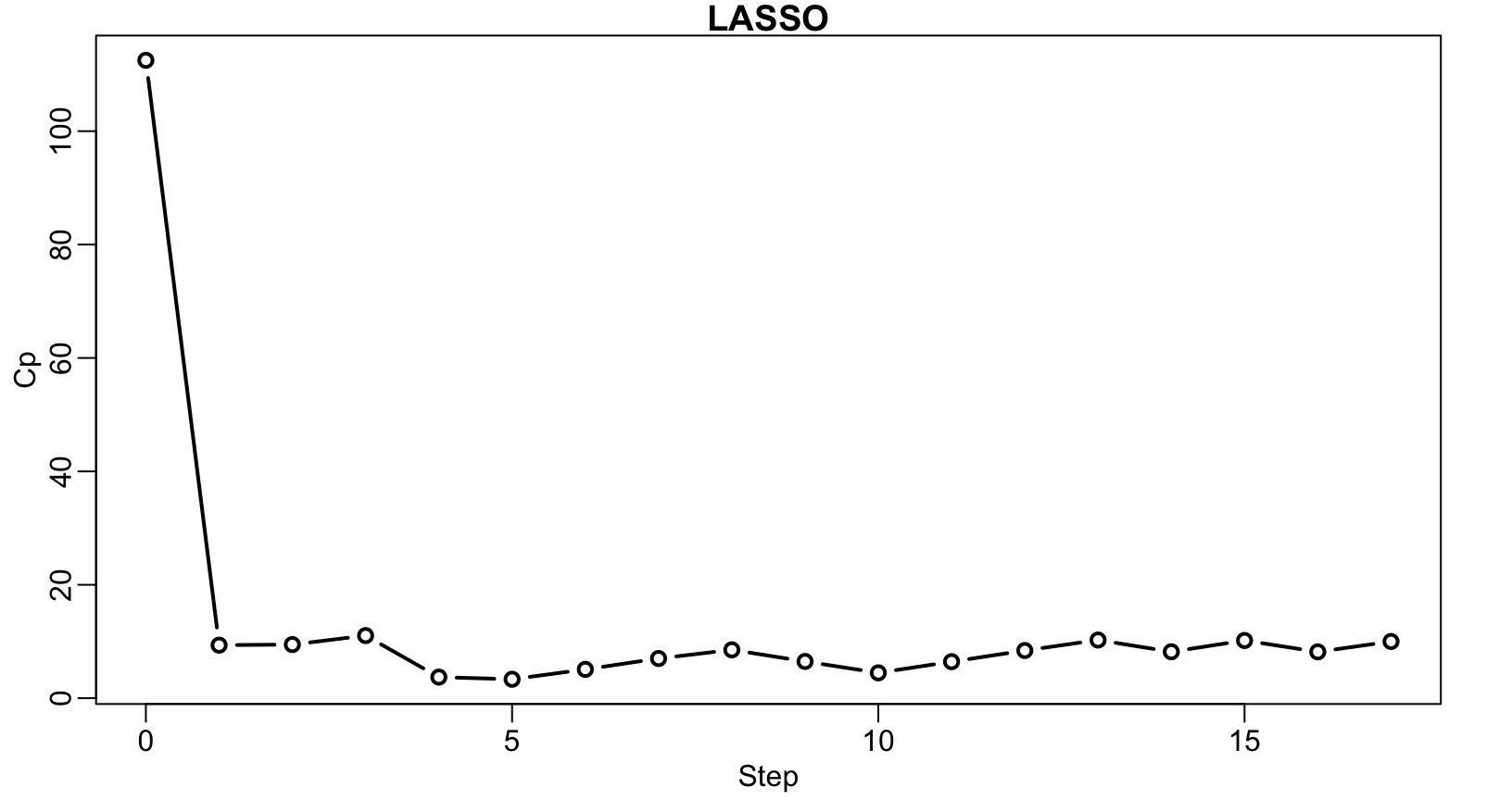
Artificial example – Lasso
## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x8 ## 8 ## [1] "x8" ## [1] "x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x6 ## 6 ## [1] "x6" "x8" ## [1] "x6+x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x1 ## 1 ## [1] "x1" "x6" "x8" ## [1] "x1+x6+x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x4 ## 4 ## [1] "x1" "x4" "x6" "x8" ## [1] "x1+x4+x6+x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x3 ## 3 ## [1] "x1" "x3" "x4" "x6" "x8" ## [1] "x1+x3+x4+x6+x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x7 ## 7 ## [1] "x1" "x3" "x4" "x6" "x7" "x8" ## [1] "x1+x3+x4+x6+x7+x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x5 ## 5 ## [1] "x1" "x3" "x4" "x5" "x6" "x7" "x8" ## [1] "x1+x3+x4+x5+x6+x7+x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x2 ## 2 ## [1] "x1" "x2" "x3" "x4" "x5" "x6" "x7" "x8" ## [1] "x1+x2+x3+x4+x5+x6+x7+x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x5 ## -5 ## [1] "x1" "x2" "x3" "x4" "x6" "x7" "x8" ## [1] "x1+x2+x3+x4+x6+x7+x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x6 ## -6 ## [1] "x1" "x2" "x3" "x4" "x7" "x8" ## [1] "x1+x2+x3+x4+x7+x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x5 ## 5 ## [1] "x1" "x2" "x3" "x4" "x5" "x7" "x8" ## [1] "x1+x2+x3+x4+x5+x7+x8" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x9 ## 9 ## [1] "x1" "x2" "x3" "x4" "x5" "x7" "x8" "x9" ## [1] "x1+x2+x3+x4+x5+x7+x8+x9" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x6 ## 6 ## [1] "x1" "x2" "x3" "x4" "x5" "x6" "x7" "x8" "x9" ## [1] "x1+x2+x3+x4+x5+x6+x7+x8+x9" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x2 ## -2 ## [1] "x1" "x3" "x4" "x5" "x6" "x7" "x8" "x9" ## [1] "x1+x3+x4+x5+x6+x7+x8+x9" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x2 ## 2 ## [1] "x1" "x2" "x3" "x4" "x5" "x6" "x7" "x8" "x9" ## [1] "x1+x2+x3+x4+x5+x6+x7+x8+x9" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x3 ## -3 ## [1] "x1" "x2" "x4" "x5" "x6" "x7" "x8" "x9" ## [1] "x1+x2+x4+x5+x6+x7+x8+x9" ## ## Call: ## lars(x = x, y = y) ## R-squared: 0.751 ## Sequence of LASSO moves: ## x8 x6 x1 x4 x3 x7 x5 x2 x5 x6 x5 x9 x6 x2 x2 x3 x3 ## Var 8 6 1 4 3 7 5 2 -5 -6 5 9 6 -2 2 -3 3 ## Step 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 ## x3 ## 3 ## [1] "x1" "x2" "x3" "x4" "x5" "x6" "x7" "x8" "x9" ## [1] "x1+x2+x3+x4+x5+x6+x7+x8+x9"
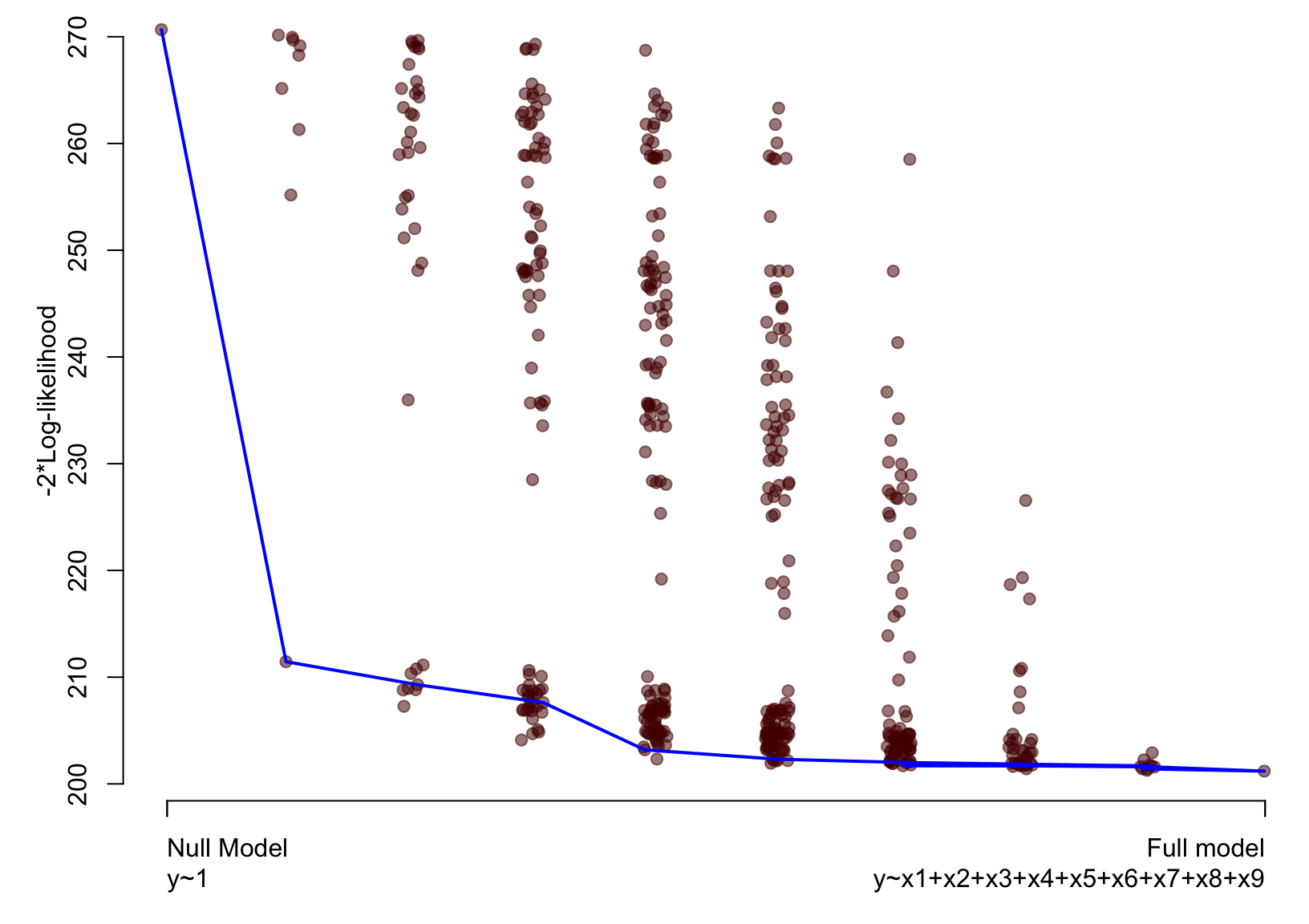
Artificial example – Lasso
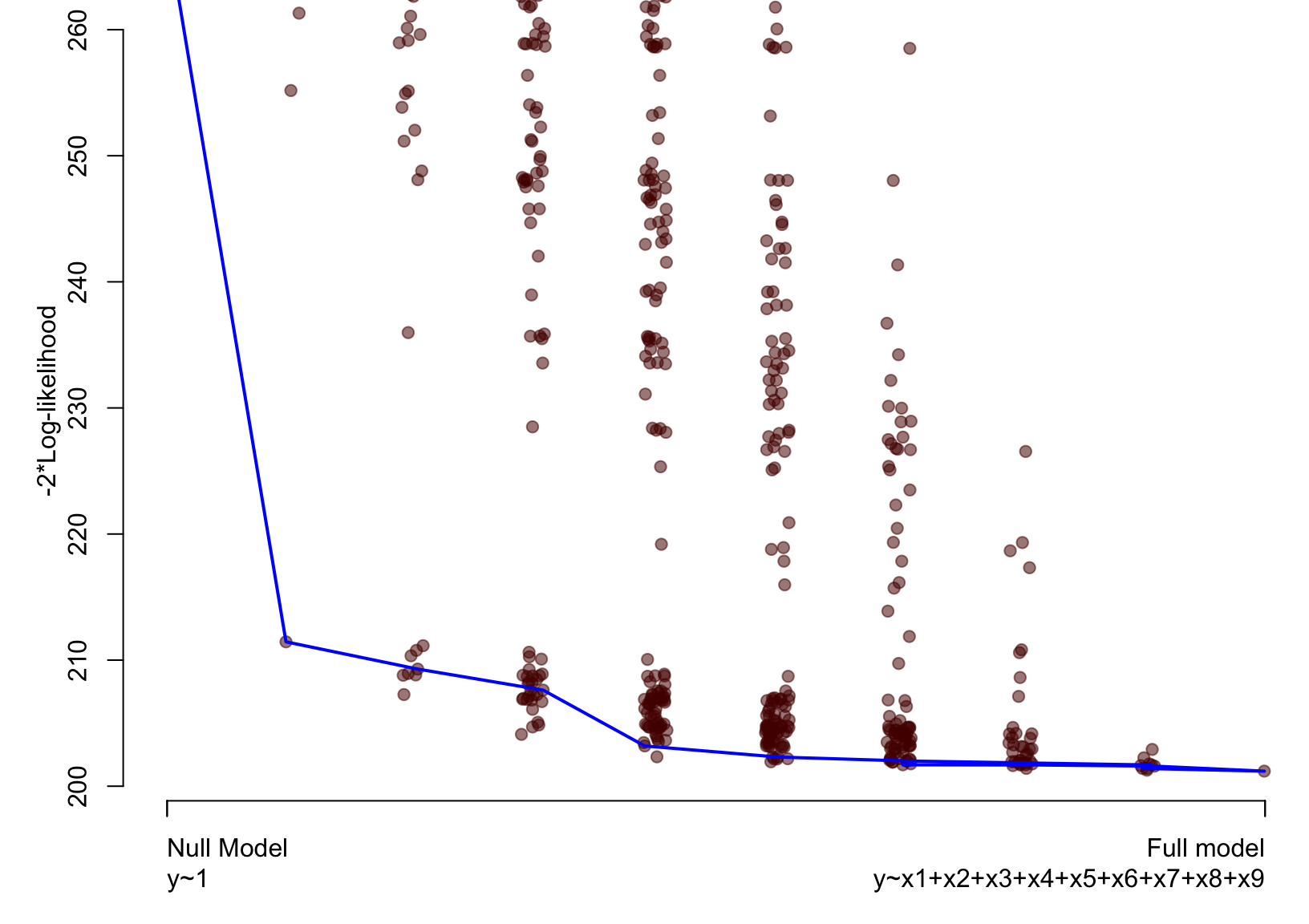
Artificial example – Lasso
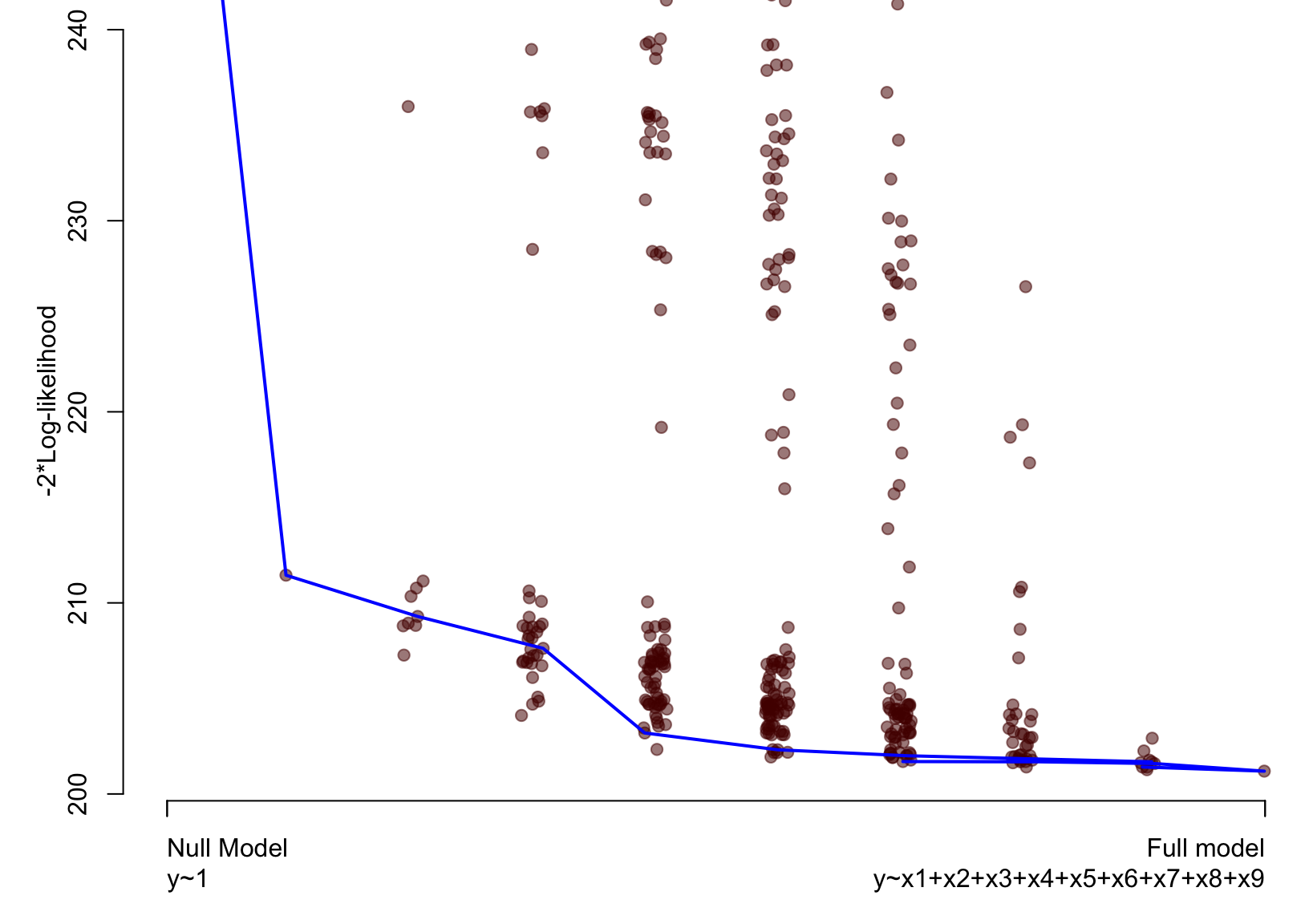
Artificial example – Lasso
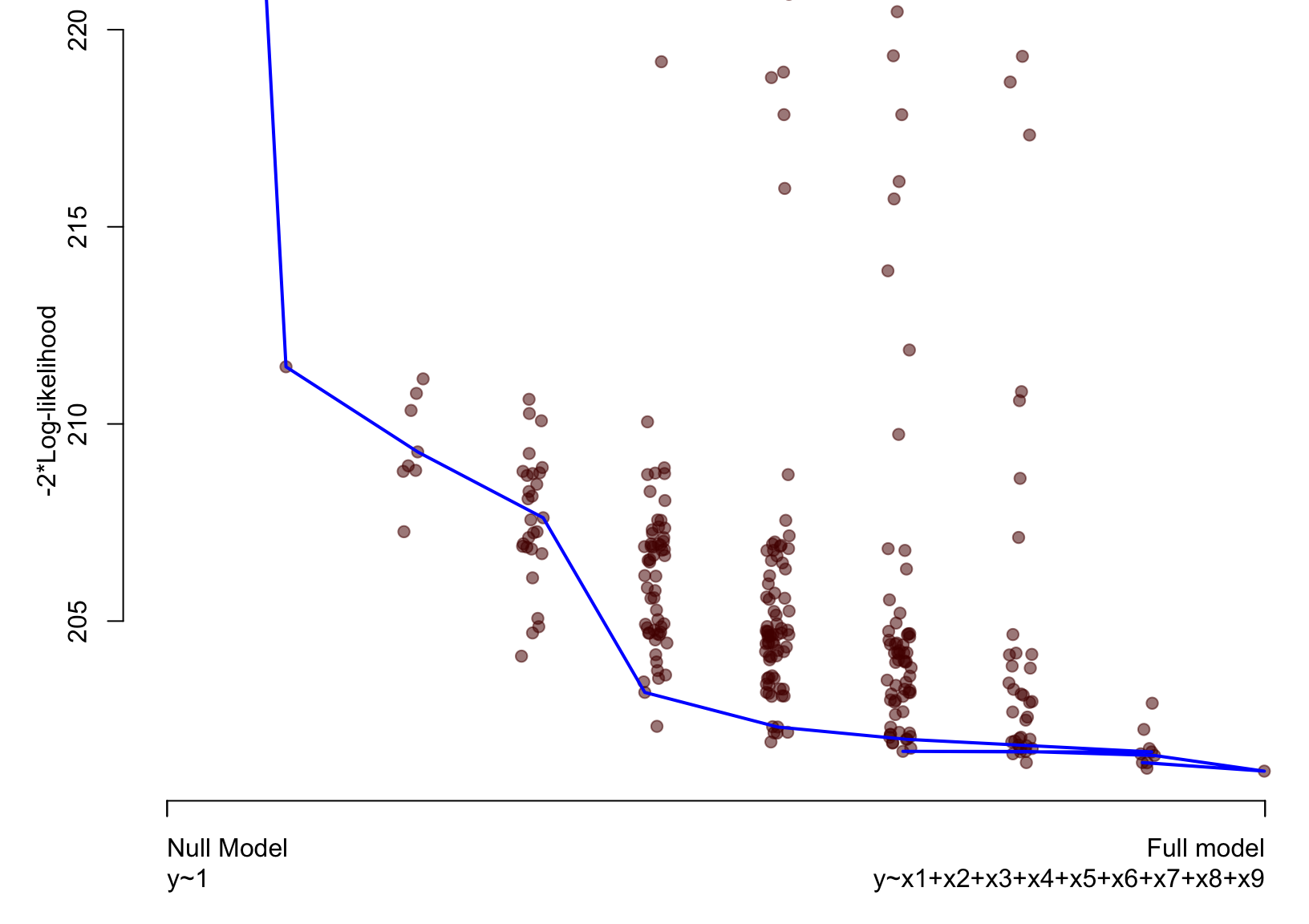
Artificial example – Lasso
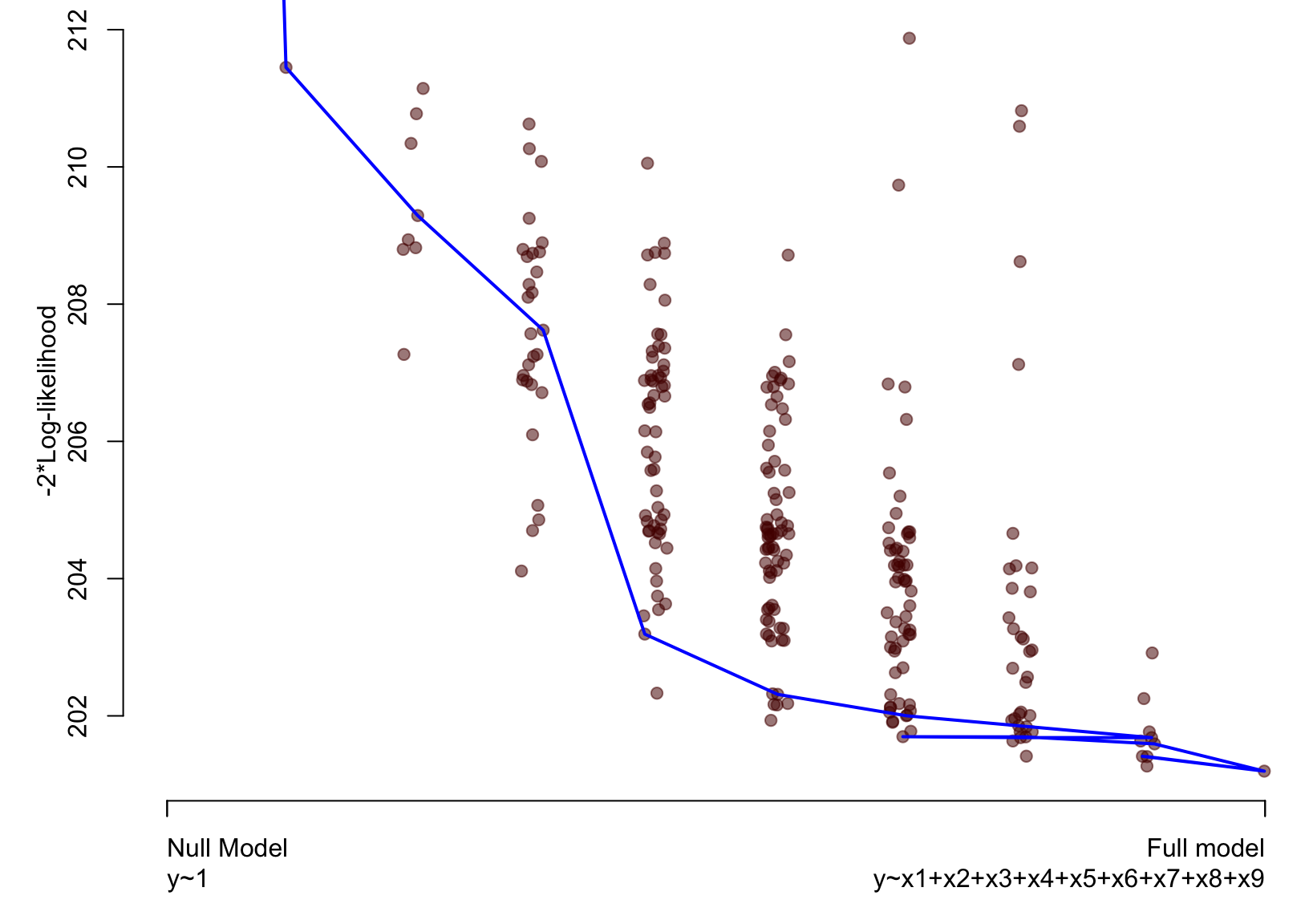
Bootstrapping the lasso
bgn.art = bglmnet(lm.art, lambda = seq(0.05, 2.75, 0.1)) plot(bgn.art, which = "boot")
Bootstrapping the lasso
bgn.art = bglmnet(lm.art) plot(bgn.art, which = "vip")
The mplot package
Get it on Github
install.packages("devtools")
require(devtools)
install_github("garthtarr/mplot",quick=TRUE)
require(mplot)
… or get it on CRAN
install.packages("mplot")
Main functions
af()for the adaptive fencevis()for VIP and model stability plotsbglmnet()bootstrapping glmnetmplot()for an interactive shiny interface
Diabetes example
Variables
| Variable | Description |
|---|---|
| age | Age |
| sex | Gender |
| bmi | Body mass index |
| map | Mean arterial pressure (average blood pressure) |
| tc | Total cholesterol (mg/dL) |
| ldl | Low-density lipoprotein ("bad" cholesterol) |
| hdl | High-density lipoprotein ("good" cholesterol) |
| tch | Blood serum measurement |
| ltg | Blood serum measurement |
| glu | Blood serum measurement (glucose?) |
| y | A quantitative measure of disease progression one year after baseline |
Diabetes data set
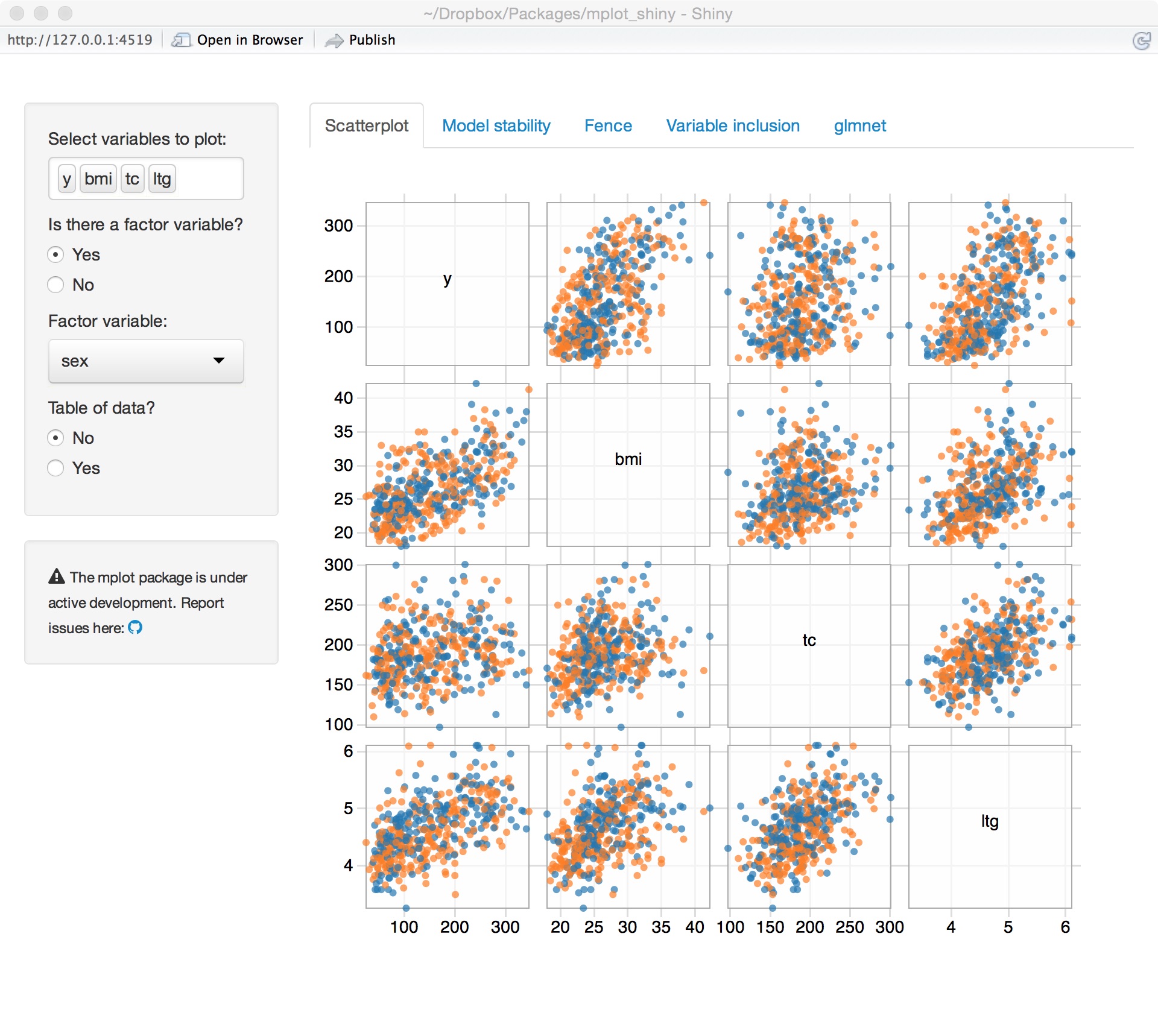
Variable importance
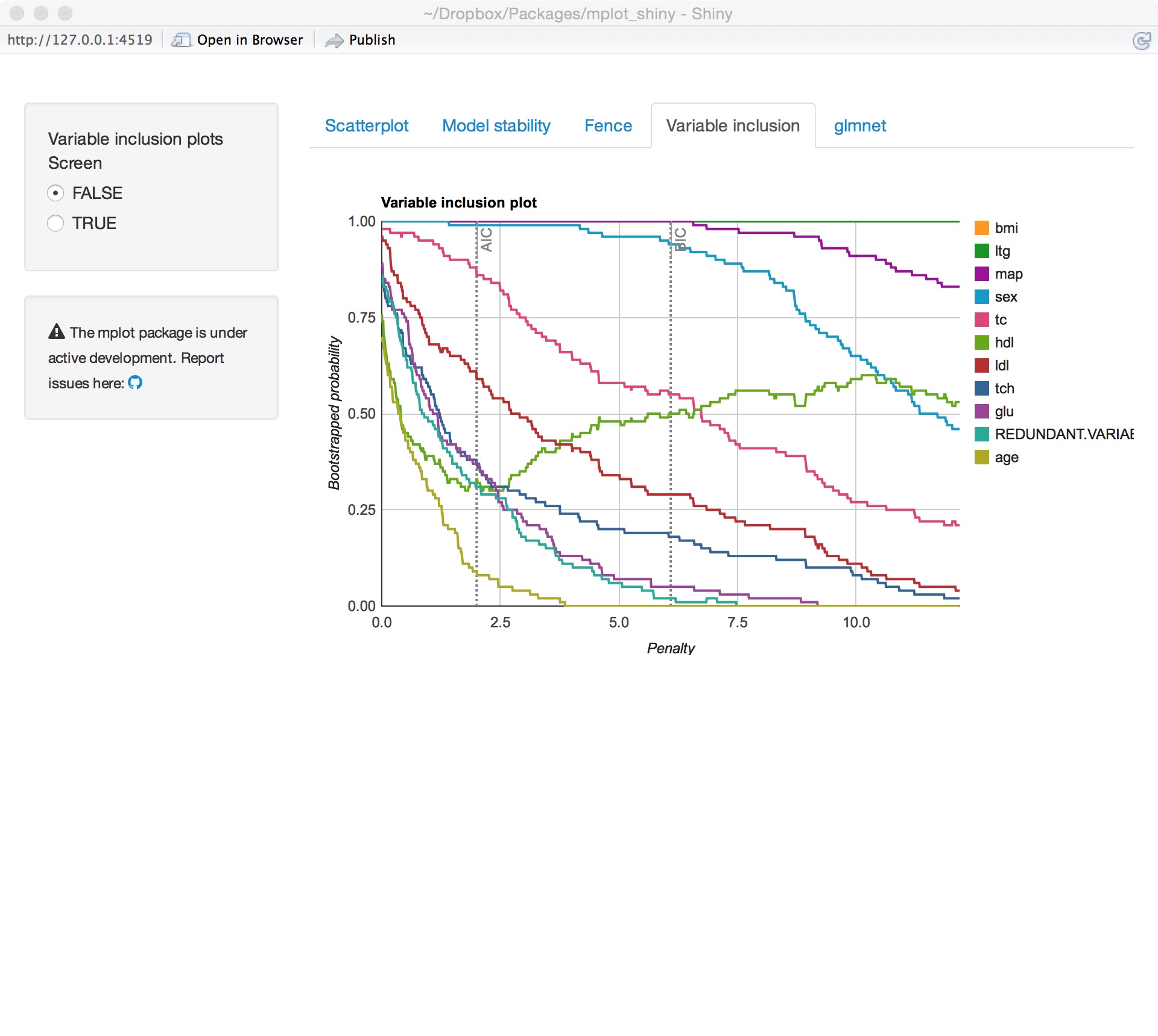
Model stability
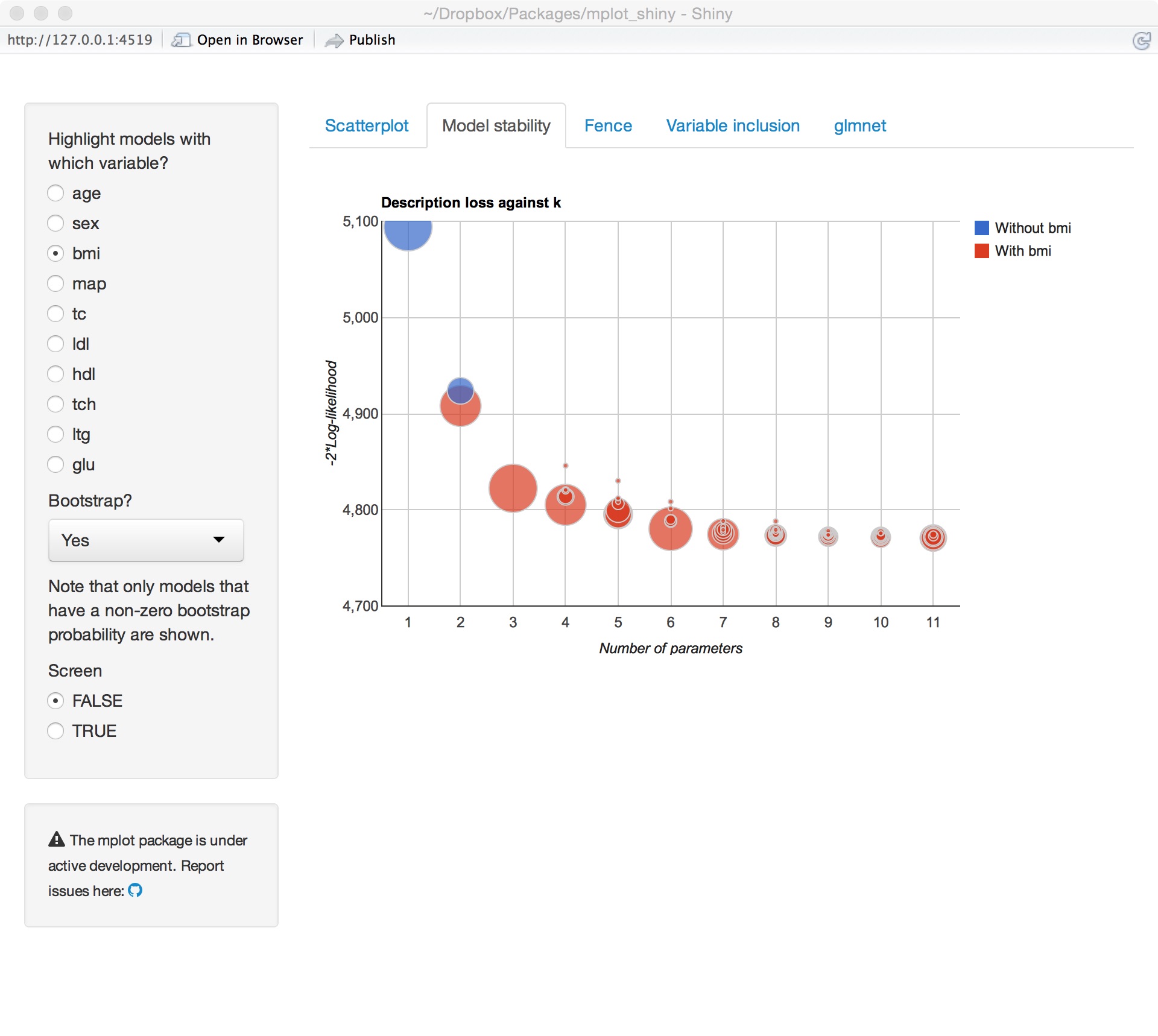
Adaptive fence
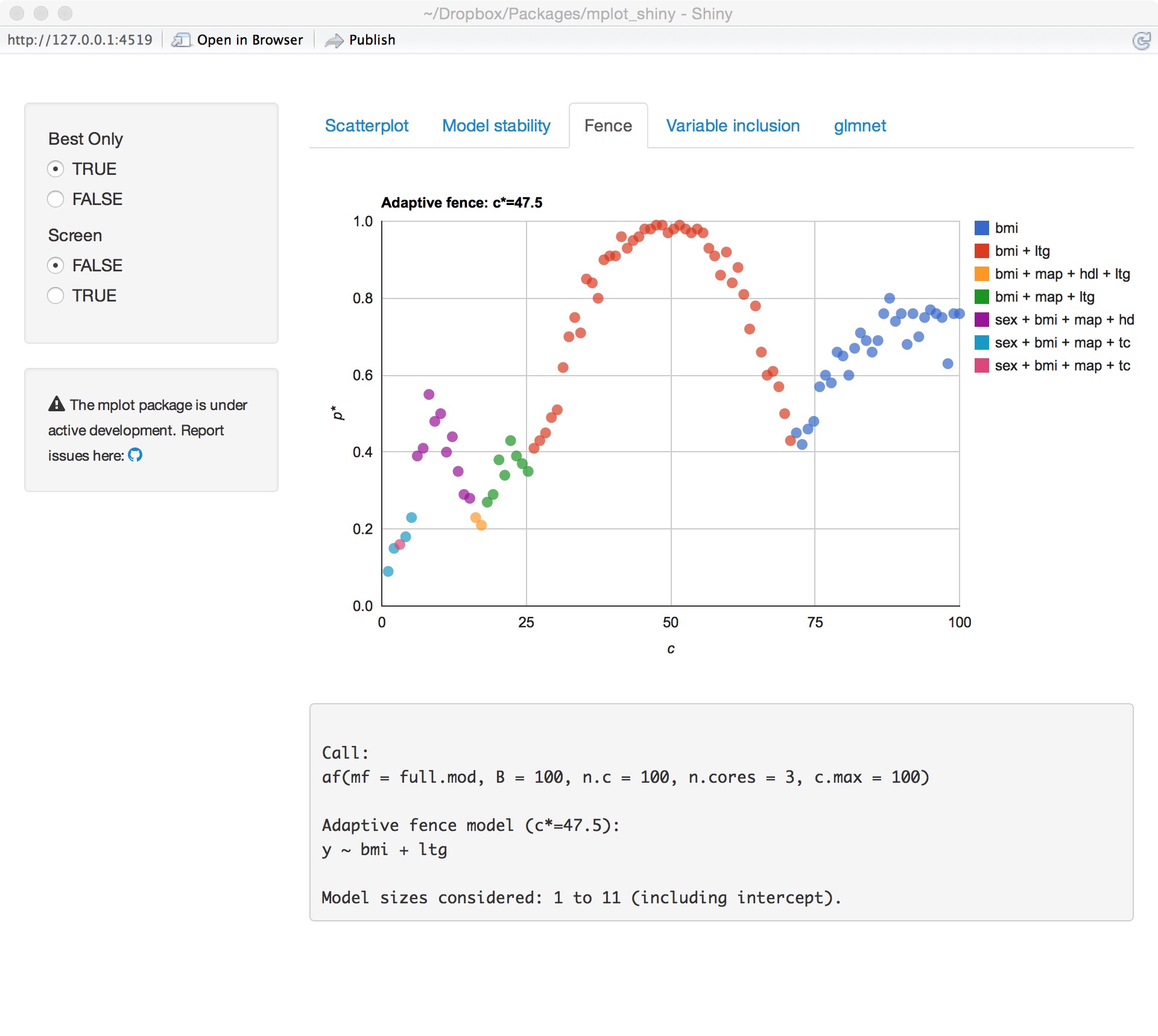
glmnet variable importance
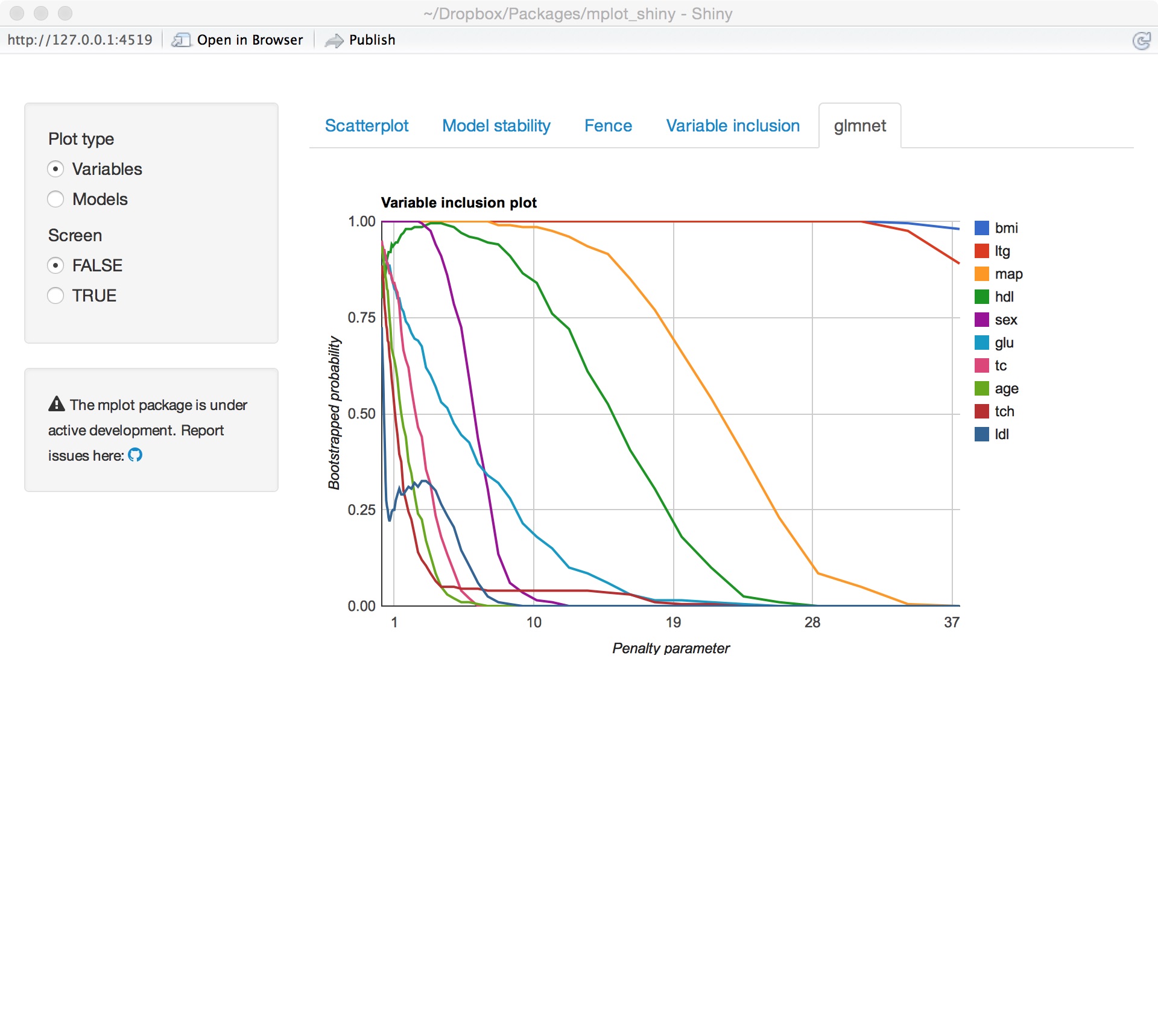
glmnet model stability
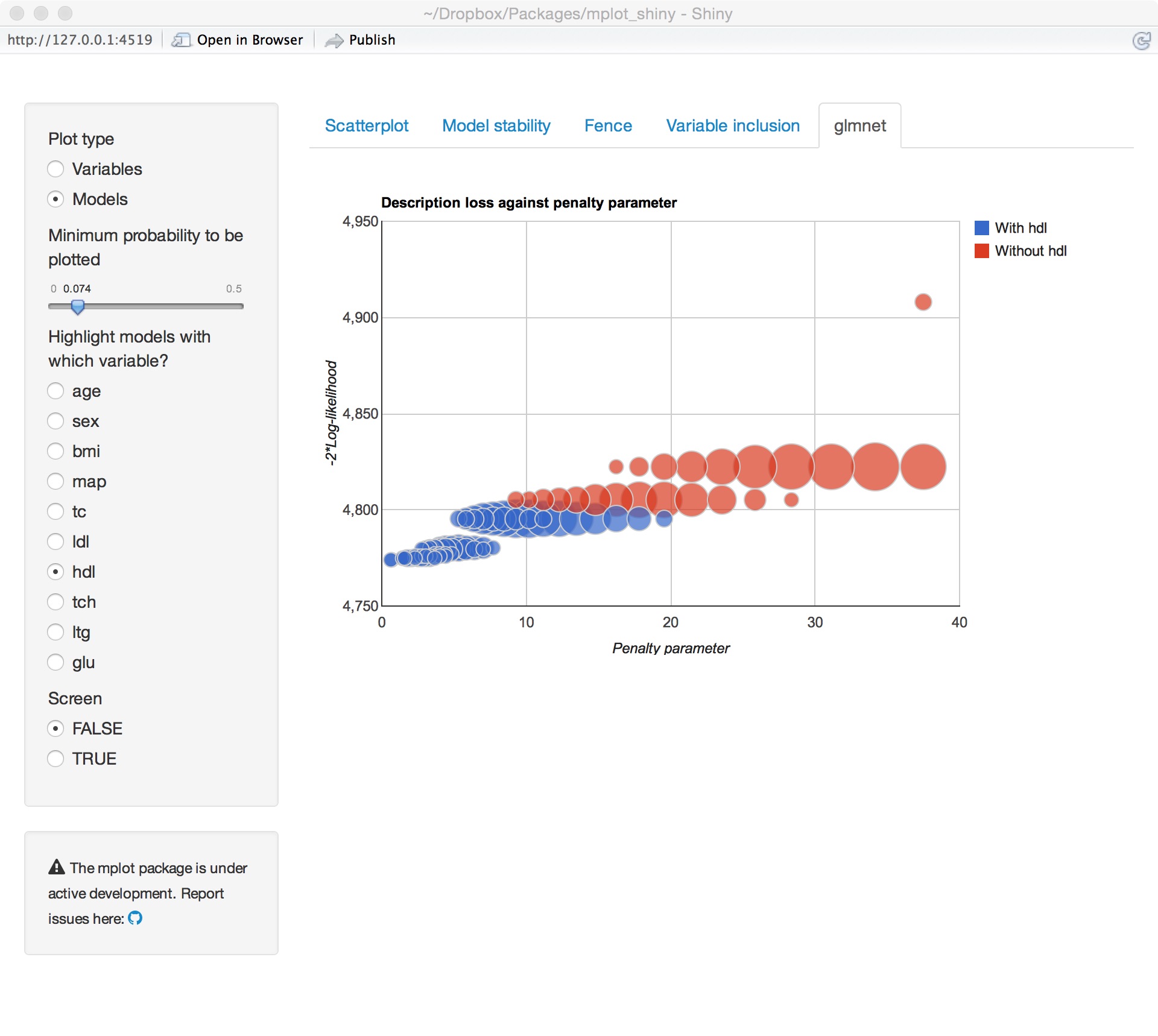
Underlying technology
R packages
- leaps
- bestglm
- googleVis
- shiny
- parallel
- mvoutlier
Web
Package development
- RStudio makes setting up package structure easy
- Version control with Github
- Documentation with
roxygen2 - Vignettes (and these slides) with markdown
devtoolsis an essential package for building and loading R packages- R packages by Hadley Wickham
Future work
Future work
- Increase the speed of GLM (approximations)?
- Mixed models (speed is an issue here too)
- Robust alternatives (other than simple screening)
- Cox regression has been requested
Find out more
- Tarr G, Mueller S and Welsh A (2015). “mplot: An R package for graphical model stability and variable selection.” arXiv:1509.07583 [stat.ME], http://arxiv.org/abs/1509.07583.
Session Info
sessionInfo()
## R version 3.2.2 (2015-08-14) ## Platform: x86_64-apple-darwin13.4.0 (64-bit) ## Running under: OS X 10.10.5 (Yosemite) ## ## locale: ## [1] en_AU.UTF-8/en_AU.UTF-8/en_AU.UTF-8/C/en_AU.UTF-8/en_AU.UTF-8 ## ## attached base packages: ## [1] stats graphics grDevices utils datasets methods base ## ## other attached packages: ## [1] lars_1.2 xtable_1.7-4 mplot_0.7.3 knitr_1.11 ## ## loaded via a namespace (and not attached): ## [1] Rcpp_0.12.1 magrittr_1.5 leaps_2.9 ## [4] doParallel_1.0.8 R6_2.1.1 foreach_1.4.2 ## [7] stringr_1.0.0 plyr_1.8.3 tools_3.2.2 ## [10] shinydashboard_0.5.1 parallel_3.2.2 htmltools_0.2.6 ## [13] iterators_1.0.7 yaml_2.1.13 digest_0.6.8 ## [16] RJSONIO_1.3-0 shiny_0.12.2 formatR_1.2.1 ## [19] codetools_0.2-14 evaluate_0.8 mime_0.4 ## [22] rmarkdown_0.8 stringi_0.5-5 compiler_3.2.2 ## [25] googleVis_0.5.10 httpuv_1.3.3
References
Efron, Bradley, Trevor Hastie, Iain Johnstone, and Robert Tibshirani. 2004. “Least Angle Regression.” The Annals of Statistics 32 (2): 407–51. doi:10.1214/009053604000000067.
Jiang, Jiming, Thuan Nguyen, and J. Sunil Rao. 2009. “A Simplified Adaptive Fence Procedure.” Statistics & Probability Letters 79 (5): 625–29. doi:10.1016/j.spl.2008.10.014.
Jiang, Jiming, J. Sunil Rao, Zhonghua Gu, and Thuan Nguyen. 2008. “Fence Methods for Mixed Model Selection.” The Annals of Statistics 36 (4): 1669–92. doi:10.1214/07-AOS517.
Meinshausen, N, and P Bühlmann. 2010. “Stability Selection.” Journal of the Royal Statistical Society: Series B (Statistical Methodology) 72 (4): 417–73. doi:10.1111/j.1467-9868.2010.00740.x.
Murray, K, S Heritier, and S Müller. 2013. “Graphical Tools for Model Selection in Generalized Linear Models.” Statistics in Medicine 32 (25): 4438–51. doi:10.1002/sim.5855.
Müller, S, and AH Welsh. 2010. “On Model Selection Curves.” International Statistical Review 78 (2): 240–56. doi:10.1111/j.1751-5823.2010.00108.x.
Tibshirani, Robert. 1996. “Regression Shrinkage and Selection via the Lasso.” Journal of the Royal Statistical Society: Series B (Methodological), 267–88.